JBrowse
The next-generation genome browser
JBrowse is a genome browser that runs on the web, on your desktop, or embedded in your app.
Check out our latest release blogpost, our embedded components, and our command line tools.
Features
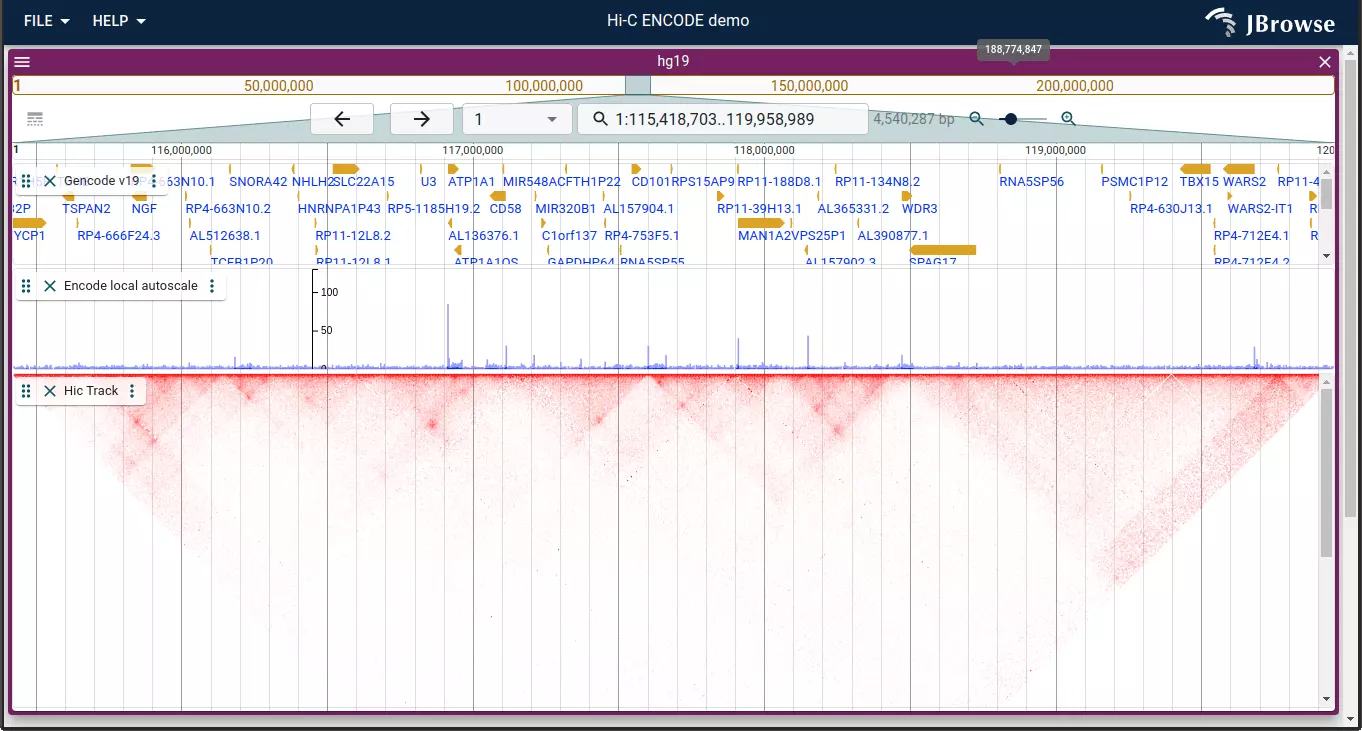
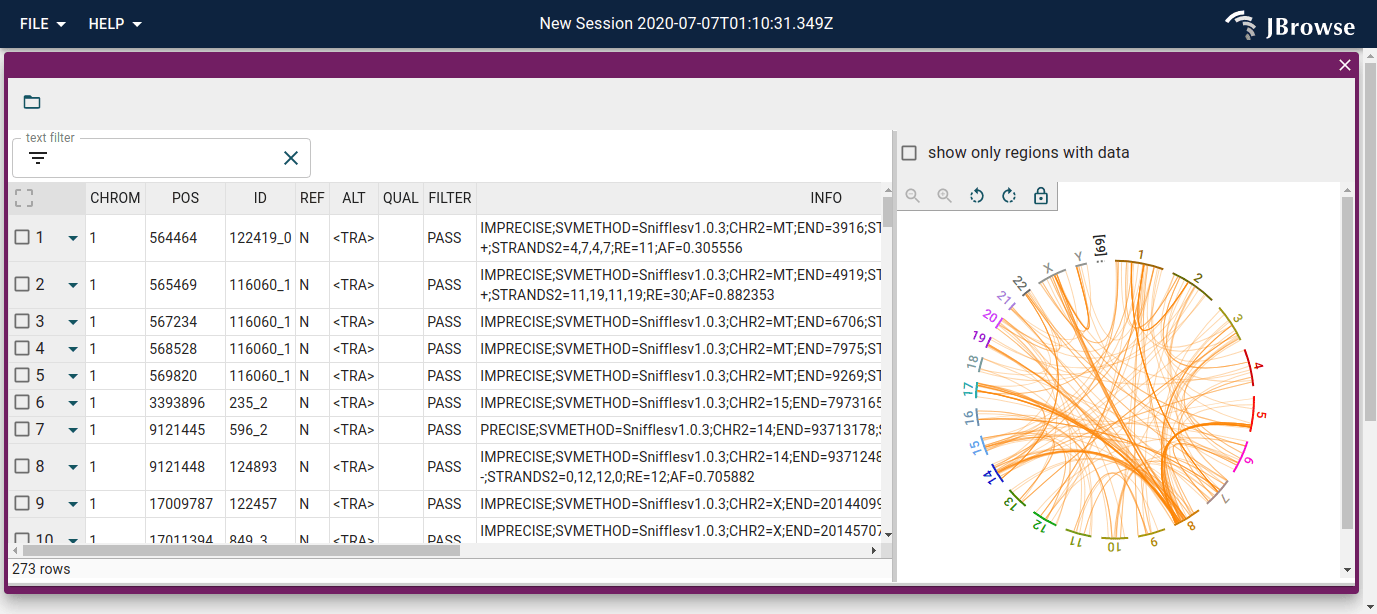
- Improved visualization of structural variants and genome synteny with tightly-integrated linear, circular, dotplot, and synteny views
- In-browser protein 3D structure displays with integrated alignment and phylogenetic tree views. Mouse over a genomic variant to see where it lands on the structure!
- Support for many common data types including BAM, CRAM, tabix indexed VCF, GFF, BED, BigBed, BigWig, and several specialized formats
- Powerful extensibility with a plugin ecosystem which can add additional view types, track types, data adapters, and more!
- See a summary of new features and a comparison to JBrowse 1
Citation
We at the JBrowse Consortium are working to make JBrowse a pluggable, open-source computational platform for integrating many kinds of biological data from many different places.
Research citations are one of the main metrics the consortium uses to demonstrate our relevance and utility when applying for funding to continue our work. If you use JBrowse in research that you publish, please cite the most recent JBrowse paper:
Diesh et al, 2023. JBrowse 2: a modular genome browser with views of synteny and structural variation. Genome Biology 24:74. https://doi.org/10.1186/s13059-023-02914-zFunding and Collaboration
JBrowse development has received support from the US National Institutes of Health (U41 HG003751), The Chan Zuckerberg Initiative, The Ontario Institute for Cancer Research (OICR), and the University of California, Berkeley.