About assemblies
What is an assembly?
An assembly in JBrowse 2 is a collection of information that describes the organism you are working with, such as its name and its reference sequence. This is what JBrowse 2 uses to set up the coordinate system for the genome browser and keep tracks organized. In JBrowse Web, you are able to have multiple assemblies on screen at the same time. JBrowse Linear Genome View, however, is more focused and allows just one assembly.
These are the options you can specify for an assembly:
- Name - The name of your assembly, often something like "GRCh38," "hg19," or "mm10".
- Aliases - Potential aliases for your assembly, such as "hg38" being an alias for "GRCh38".
- Reference Sequence - The file that has the genome sequence for your organism. JBrowse natively supports indexed FASTA (compressed or not), 2BIT, and chrom.sizes sequence files.
- Reference Sequence Name Aliases - You can define aliases for the names in your reference sequence file, so if for example your reference sequence uses "chr1", you can add an alias for it so that data file that use "1" instead will still display correctly.
- Colors - You can define a list of colors that will be used to color the reference sequences. If you supply fewer colors than reference sequences, JBrowse will cycle through them. By default JBrowse will use its built-in color scheme of 26 colors based on the UCSC Genome Browser color scheme.
Adding an assembly
An assembly definition is a JSON object that describes the assembly. You can create them by hand, but an easier way to do it is to generate them via the JBrowse command-line interface (CLI), so we'll use that to generate our assembly.
Installing JBrowse CLI
The JBrowse 2 CLI is designed to set up a configuration file for JBrowse Web, but since JBrowse Linear Genome View is based on the same code, we can use it to generate what we need as well.
The instructions for installing JBrowse CLI are on the
"Introduction" page. After installing, if you run the
command jbrowse --help in the terminal, you should see something like this:
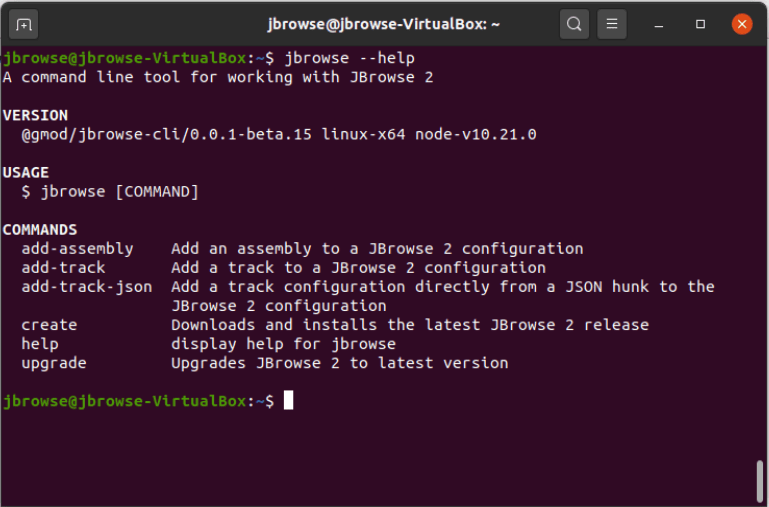
If you're having trouble installing the JBrowse CLI globally, or if you don't
want to install it, you can also do all of the things we will do in this
tutorial by replacing jbrowse in any commands with npx @jbrowse/cli. For
example, to get the help output you would run
sh npx @jbrowse/cli --help
Using this command will create a temporary copy of the JBrowse CLI that doesn't get installed.
Gathering files
Let's use a human GRCh38 assembly as our example. First we need a sequence file. We'll use this url: https://s3.amazonaws.com/jbrowse.org/genomes/GRCh38/fasta/GRCh38.fa.gz.
We also need to define some reference sequence name aliases so that any files we want to use that use "chr1" instead of "1" will display correctly. There is an alias file that we can use at http://s3.amazonaws.com/jbrowse.org/genomes/GRCh38/GRCh38.aliases.txt.
This aliases file has on each line a name from the reference sequence and then a tab-separated list of aliases for that name.
Running the jbrowse command
We can use JBrowse CLI to create a JBrowse Web config file, which we can use to
get the assembly for JBrowse Linear Genome View. We create a config file with an
assembly (or add an assembly to an existing config file) with the
jbrowse add-assembly command. Go ahead and run jbrowse add-assembly --help
to see all the options available.
There are a lot of options, and not all of them are things we need to worry about in this case. Now go ahead and run the below command, referring to the help to interpret the options as needed:
jbrowse add-assembly https://s3.amazonaws.com/jbrowse.org/genomes/GRCh38/fasta/GRCh38.fa.gz \
--name GRCh38 \
--alias hg38 \
--refNameAliases http://s3.amazonaws.com/jbrowse.org/genomes/GRCh38/GRCh38.aliases.txt \
--skipCheck
This will create a file called config.json. Go ahead and open that file up. In
it, you will see an entry called "assemblies". The first (and only) entry in
that list is our assembly. You can see the name, alias, sequence, and reference
name aliases we specified in our command. If for some reason the index files
were unusually named, you could change them in the "faiLocation" and
"gziLocation" entries, but these look right. Create a new file called
"assembly.js" and copy the assembly entry over to that file and have it be
exported, like this:
export default {
name: 'GRCh38',
sequence: {
type: 'ReferenceSequenceTrack',
trackId: 'GRCh38-ReferenceSequenceTrack',
adapter: {
type: 'BgzipFastaAdapter',
fastaLocation: {
uri: 'https://s3.amazonaws.com/jbrowse.org/genomes/GRCh38/fasta/GRCh38.fa.gz',
locationType: 'UriLocation',
},
faiLocation: {
uri: 'https://s3.amazonaws.com/jbrowse.org/genomes/GRCh38/fasta/GRCh38.fa.gz.fai',
locationType: 'UriLocation',
},
gziLocation: {
uri: 'https://s3.amazonaws.com/jbrowse.org/genomes/GRCh38/fasta/GRCh38.fa.gz.gzi',
locationType: 'UriLocation',
},
},
},
aliases: ['hg38'],
refNameAliases: {
adapter: {
type: 'RefNameAliasAdapter',
location: {
uri: 'http://s3.amazonaws.com/jbrowse.org/genomes/GRCh38/GRCh38.aliases.txt',
locationType: 'UriLocation',
},
},
},
}
Then add an import to your "index.html" adding this data:
<html>
<head>
<script
src="//unpkg.com/react@16/umd/react.development.js"
crossorigin
></script>
<script
src="//unpkg.com/react-dom@16/umd/react-dom.development.js"
crossorigin
></script>
<script
src="//unpkg.com/@jbrowse/react-linear-genome-view/dist/react-linear-genome-view.umd.development.js"
crossorigin
></script>
</head>
<body>
<h1>We're using JBrowse Linear Genome View!</h1>
<div id="jbrowse_linear_genome_view"></div>
<script type="module">
import assembly from './assembly.js'
</script>
</body>
</html>
Keep the config.json around for now, as we'll use it in the next step as well.