JBrowse web setup using the CLI
Prerequisites
- Node.js 18+. Note: we recommend not using
aptto install Node.js, it often installs old versions. Good alternatives include NodeSource or NVM. - Samtools installed e.g.
sudo apt install samtoolsorbrew install samtools, used for creating FASTA indexes and BAM/CRAM index files - tabix installed e.g.
sudo apt install tabixorbrew install htslib, used for creating tabix indexes for BED/VCF/GFF files
Installing the JBrowse CLI
The JBrowse CLI can perform many tasks to help you manage JBrowse 2, such as:
- create a new instance of JBrowse 2 automatically
- update an existing instance of JBrowse 2 with the latest released version
- configure your JBrowse 2 instance
To globally install the JBrowse CLI, run
npm install -g @jbrowse/cli
Test the installation with:
jbrowse --version
If you can't or don't want to globally install the JBrowse CLI, you can also use
the npx command, which
is included with Node.js, to run JBrowse CLI without installing it. Simply
replace jbrowse with npx @jbrowse/cli in any command, e.g.
npx @jbrowse/cli --version
Alternatively, download a single-file bundle:
wget https://unpkg.com/@jbrowse/cli/bundle/index.js -O jbrowse
chmod +x jbrowse
./jbrowse --help
Using jbrowse create to download JBrowse 2
In the directory where you would like to download JBrowse 2, run
jbrowse create jbrowse2
This downloads and unzips jbrowse-web into a folder named "jbrowse2" (the name is arbitrary — it's just static html/css/js files).
Alternatively, you can download the zip manually from https://github.com/GMOD/jbrowse-components/releases
Checking the download
The directory where you downloaded JBrowse should look something like this:
jbrowse2/
- asset-manifest.json
- favicon.ico
- index.html
- manifest.json
- robots.txt
- static/
- test_data/
- version.txt
Running JBrowse 2
JBrowse 2 requires a web server to run. It won't work if you try to directly
open the index.html in your web browser.
Typically you place the folder in a web server's static directory (e.g.
/var/www/html/jbrowse2/) and visit http://yourserver/jbrowse2.
To quickly verify the download locally, run a development server:
cd jbrowse2/
npx serve .
# or
npx serve -S . # if you want to refer to symlinked data later on
Navigate to the location shown in the CLI output (likely
http://localhost:3000).
Your page should look something like this:
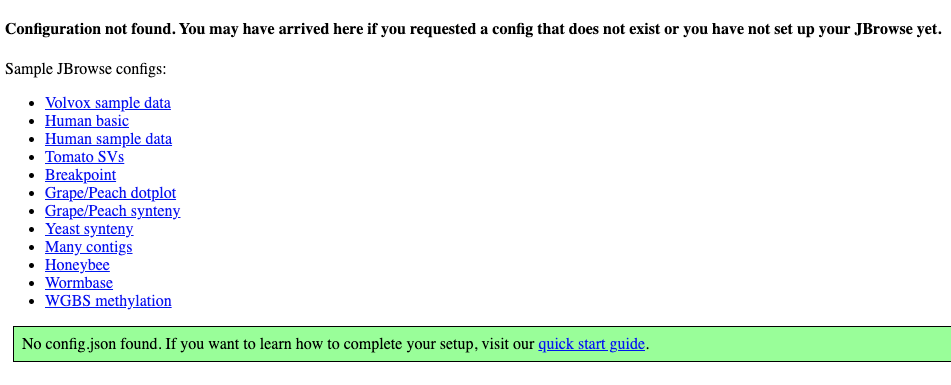
Click on the sample config to see JBrowse 2 running with a demo configuration. It should look like this:
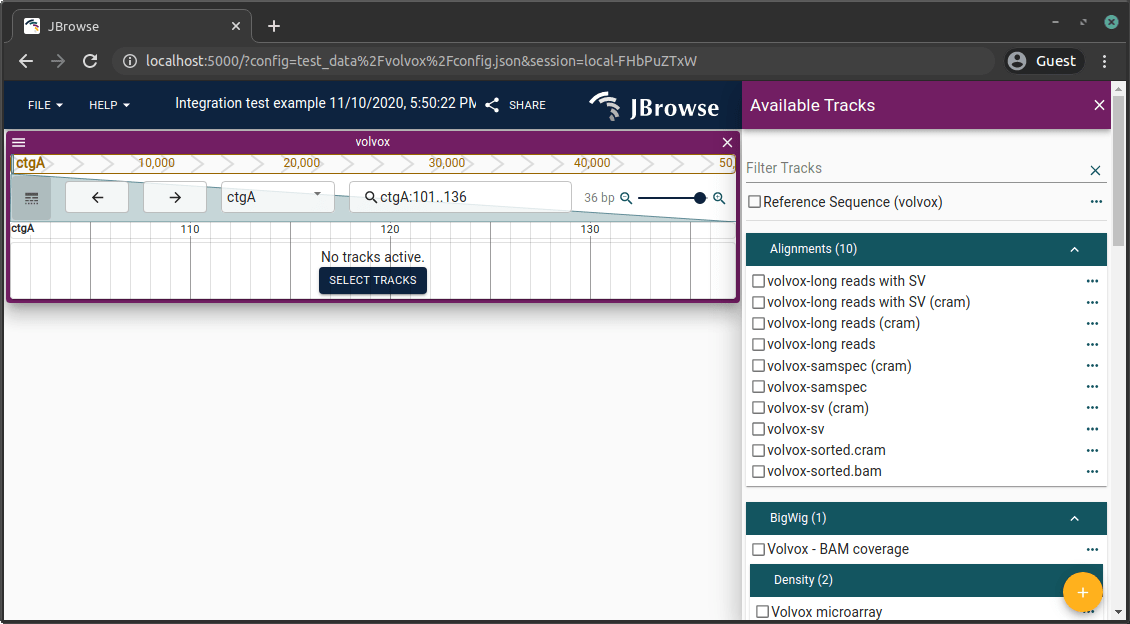
Congratulations! You're running JBrowse 2.
Adding tracks
Adding a genome assembly in FASTA format
The first step is to index your FASTA with samtools, then add the assembly:
samtools faidx genome.fa
jbrowse add-assembly genome.fa --load copy --out /var/www/html/jbrowse/
This writes an assembly entry to /var/www/html/jbrowse/config.json (creating
it if needed) and copies genome.fa and genome.fa.fai into that directory.
Use --load symlink to symlink the files instead of copying.
JBrowse 2 also supports other assembly file formats, including bgzip-compressed indexed FASTA, and 2bit files.
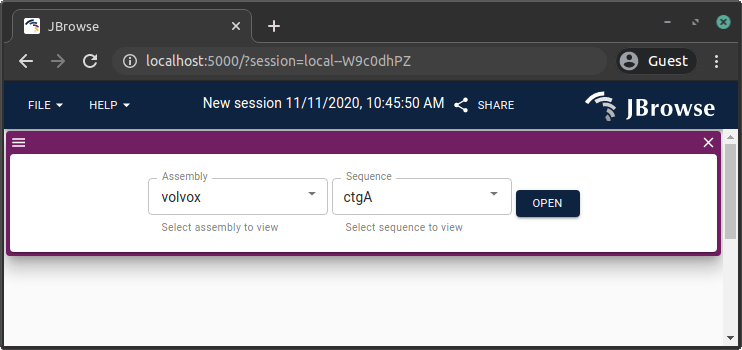
Adding a BAM or CRAM track
samtools index file.bam
jbrowse add-track file.bam --load copy --out /var/www/html/jbrowse
samtools index file.cram
jbrowse add-track file.cram --load copy --out /var/www/html/jbrowse
This adds a track entry to config.json and copies the files. Use
--load symlink to symlink instead. Run jbrowse add-track --help for more
options.
After adding a track, refresh the page, open a linear genome view, then open the track selector to see the new track.
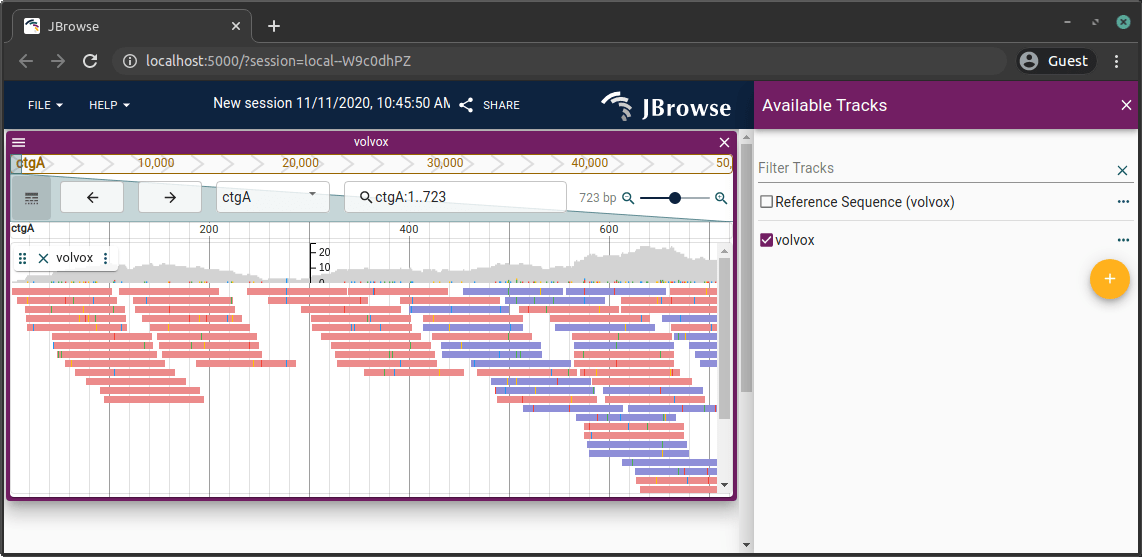
Adding a VCF track
VCFs must be bgzip-compressed and tabix-indexed:
bgzip file.vcf
tabix file.vcf.gz
jbrowse add-track file.vcf.gz --load copy --out /var/www/html/jbrowse
If tabix reports that your VCF is not sorted, sort it first with bcftools:
bcftools sort file.vcf > file.sorted.vcf
bgzip file.sorted.vcf
tabix file.sorted.vcf.gz
You can also bgzip and index with the bcftools tool.
bcftools view volvox.vcf --output-type z > volvox.vcf.gz
rm volvox.vcf
bcftools index --tbi volvox.vcf.gz
For more info about bgzip, tabix, and bcftools, see
https://www.htslib.org/.
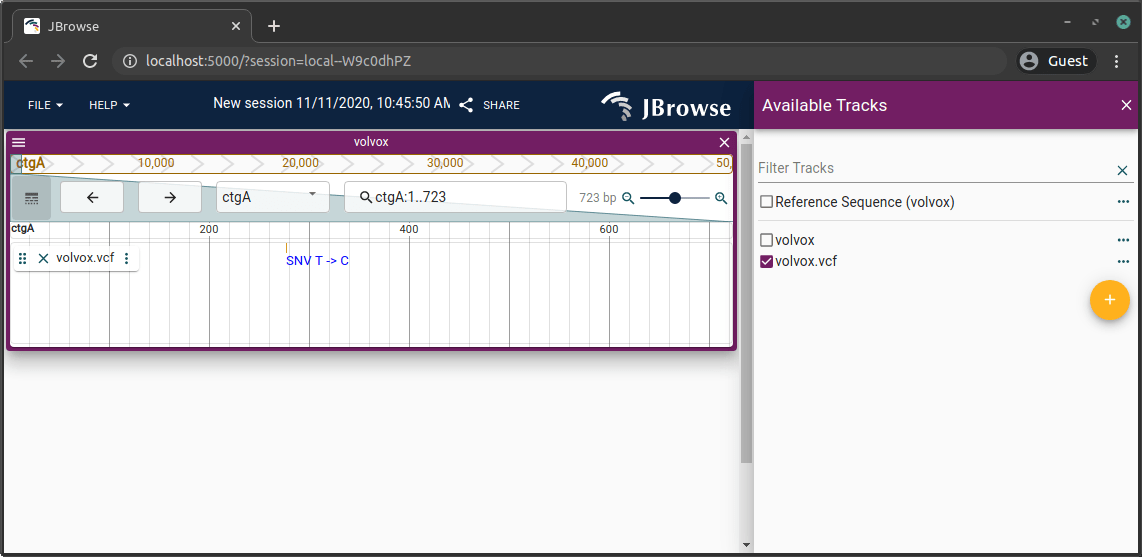
Adding a BigWig/BigBed track
BigWig/BigBed files require no external index, so loading is straightforward:
jbrowse add-track file.bw --load copy --out /var/www/html/jbrowse
Adding a GFF3 file with GFF3Tabix
jbrowse sort-gff yourfile.gff | bgzip > yourfile.sorted.gff.gz
tabix yourfile.sorted.gff.gz
jbrowse add-track yourfile.sorted.gff.gz --load copy
The jbrowse sort-gff command is equivalent to (from the
tabix documentation):
(grep "^#" in.gff; grep -v "^#" in.gff | sort -t"`printf '\t'`" -k1,1 -k4,4n) > sorted.gff;
Adding a synteny track from a PAF file
Here we use the grape and peach genome assemblies, but replace with your own data if applicable.
Use minimap2 to create a PAF file from FASTA files:
## Use minimap2 to create a PAF from your assemblies
## -cx asm20 is appropriate for cross-species comparisons (~5% divergence)
## use asm5 for same-species, asm10 for moderately diverged strains
## consult the minimap2 docs or published protocols for your organisms
minimap2 -cx asm20 grape.fa peach.fa > peach_vs_grape.paf
## add each assembly to jbrowse config
## the -n flag names the assemblies explicitly
jbrowse add-assembly grape.fa --load copy -n grape --out /var/www/html/jbrowse
jbrowse add-assembly peach.fa --load copy -n peach --out /var/www/html/jbrowse
Next, we'll load the synteny "track" from the PAF file.
Order matters here for the --assemblyNames parameter:
If minimap2 is run as minimap2 grape.fa peach.fa, then you need to load as
--assemblyNames peach,grape.
The order is reversed between the minimap2 and jbrowse tools.
jbrowse add-track peach_vs_grape.paf --assemblyNames peach,grape --load copy --out /var/www/html/jbrowse
Indexing feature names for searching
Optionally, build a text index so users can search by gene name or feature ID:
jbrowse text-index --out /var/www/html/jbrowse
This indexes Gff3Tabix tracks (gene names/IDs) and VcfTabix tracks (variant IDs), printing a progress bar per track. Once complete, gene names can be typed directly into the LGV search box.
See the text-index command docs for more info. Also see the FAQ entries for text searching
Conclusion
With an assembly and tracks configured, you're ready to explore your data. Useful next steps:
- User guide — navigating views, track types, and UI features
- Config guide — advanced track and assembly configuration
- JBrowse CLI reference — full reference for all CLI commands used here
- FAQ — common questions including text searching and CORS issues
Miscellaneous tips
You can use --subDir to organize your data directory:
mkdir my_bams
## Copies .bam and .bai files to my_bams folder
jbrowse add-track myfile.bam --subDir my_bams --load copy --out /var/www/html/jbrowse
Upgrade JBrowse Web to the latest version
You can upgrade your JBrowse release to the latest version with:
# run this command in an existing jbrowse 2 installation
jbrowse upgrade
This downloads the latest jbrowse-web release and unzips it into the current directory.
Upgrade @jbrowse/cli to the latest
To upgrade the CLI tools, you can re-run the install command
npm install -g @jbrowse/cli
Output to a custom config file name
You can use filenames other than config.json, and put configs in subfolders of your jbrowse 2 installation too
jbrowse add-assembly mygenome.fa --out /path/to/my/jbrowse2/alt_config.json --load copy
This would then be accessible at e.g. http://localhost/jbrowse2/?config=subfolder/alt_config.json