v1.6.6 Release
We are pleased to release v1.6.6!
This contains several updated synteny features, including:
- load .chain files from UCSC
- load .delta files from mummer
- load .anchors and .anchors.simple files from MCScan from the GUI
- ability to "rectangularize" the dotplot view and improved dotplot view overviews (shows total bp of selected regions)

Screenshot showing updated import form with ability to load MCScan data

Screenshot showing multiple tracks open at once, with the .anchors (green, gene pairs) and .anchors.simple (black, larger synteny blocks) files from MCScan shown in the dotplot view
Note: MCScan functionality existed internally but was not easy to use. The configuration schema for MCScan anchors files changed also to load the .bed files at startup. See the config guide for details
We also have an updated configuration editor GUI with collapsible sections
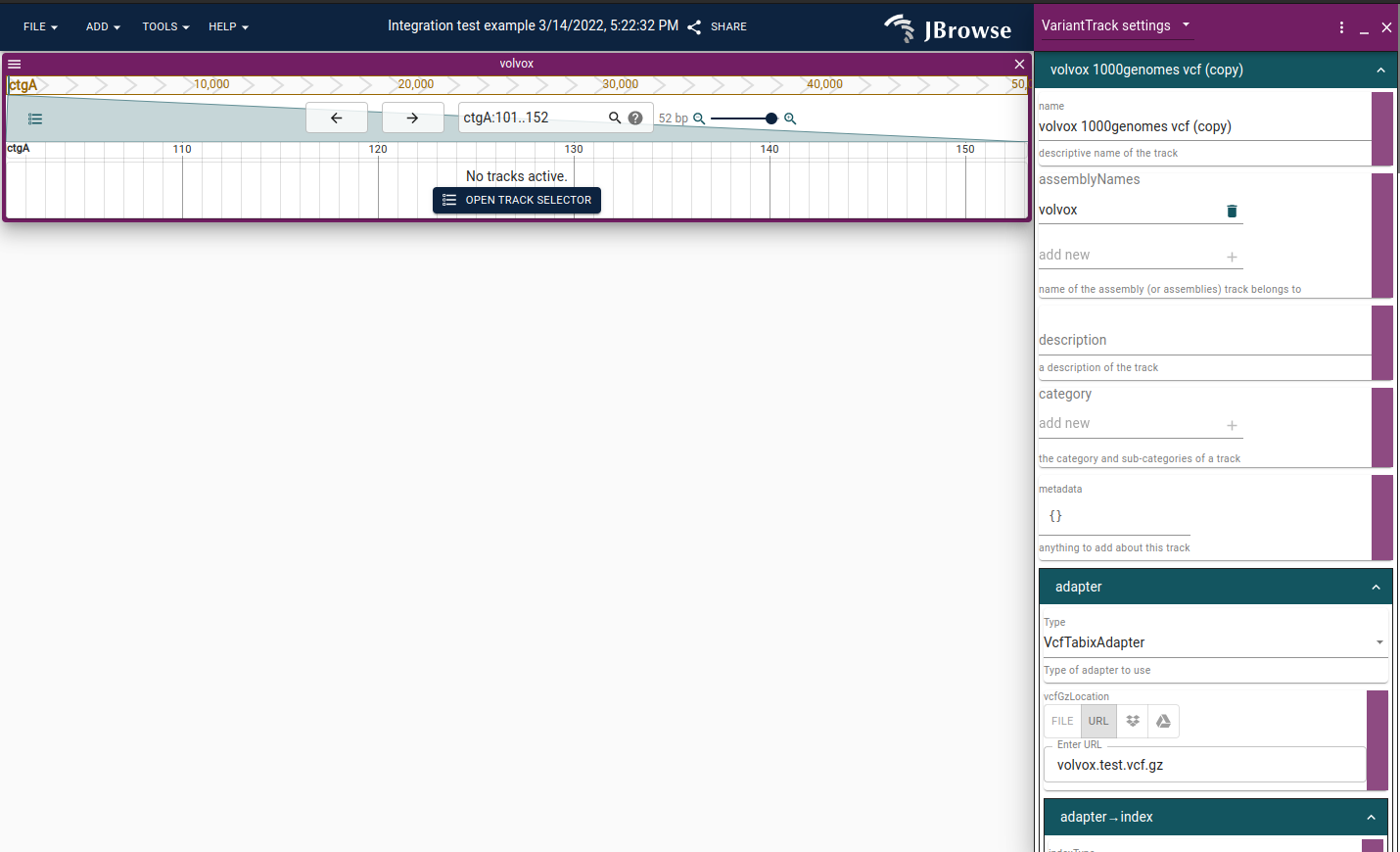
Screenshot showing new collapsible (turquoise) sections in the config editor
A number of alignments track look and feel improvements were also added. The scale bar was made to just list the min and max values when the height is small, clip indicator position when horizontally flipped was fixed, and the headroom was removed making the snpcoverage take up the full height of the small area it uses.

There have also been some changes to how internet accounts work. If you use one of the built-in accounts like Google Drive or Dropbox, there's no need to change anything. If you've implemented your own internet account, though, see #2725 for more details.
Downloads
To install JBrowse 2 for the web, you can download the link above, or you can use the JBrowse CLI to automatically download the latest version. See the JBrowse web quick start for more details.
1.6.6 (2022-03-15)
🚀 Enhancement
core- #2796 Add collapsible accordion sections in configuration editor (@cmdcolin)
- #2791 Add new coloring options for dotplot and ability to "rectangularize" dotplot view (@cmdcolin)
- #2741 Allow ability to enter a space-separated locstring to open a list of regions (@cmdcolin)
- #2725 Refactor InternetAccounts, add standard getFetcher (@garrettjstevens)
- #2787 Display the total bp viewed in the header of the dotplot view (@cmdcolin)
- #2767 Wiggle and SNPCoverage look and feel improvements (@cmdcolin)
- #2746 Add .delta and .chain format adapters, fix ref name aliasing in synteny/dotplot views, and optimize very long CIGAR string in synteny view (@cmdcolin)
- Other
- #2799 Exit process after rendering to speed up jb2export (@cmdcolin)
- #2793 Add abortcontroller polyfill to jbrowse-img to allow it to run under node 14 (@cmdcolin)
- #2761 Add a --clean
argument to
jbrowse upgradeto clean up old files (@cmdcolin) - #2760 Make a configurable refNameColumn in RefNameAliasAdapter (@cmdcolin)
🐛 Bug Fix
core- #2798 Fix bug where web worker would sometimes be called before it was ready (@garrettjstevens)
- Other
- #2797 Fix crash plotting methylation in sparse regions (@cmdcolin)
- #2782 Fix display of cytobands when horizontally flipped (@cmdcolin)
- #2678 Preserve double border line when using trackLabel offset and use smaller gap between snpcoverage and reads (@cmdcolin)
- #2774 Fix overwriting broken symlink with --force in add-track CLI (@cmdcolin)
- #2773 Fix using global stats autoscale on wiggle tracks (@cmdcolin)
- #2766 Add a check for empty content blocks to fix rare empty stats estimation (@cmdcolin)
📝 Documentation
- #2804 Add note about additional pre-requisites to README (@cmdcolin)
- #2762 Add bookmark widget docs to user guide (@cmdcolin)
🏠 Internal
- #2813 Create codeVerifierPKCE only when needed (@garrettjstevens)
- #2808 Polyfill window.crypto.getRandomValues in tests (@garrettjstevens)
Committers: 2
- Colin Diesh (@cmdcolin)
- Garrett Stevens (@garrettjstevens)