v1.3.0 Release
We're excited to announce the v1.3.0 release of JBrowse Web! Some highlights of this release include:
In-app plugin store
Building on the plugin store on our website in the last release, we're now excited to announce that plugins can be installed from within JBrowse Web! Plugins from our plugin store can now be installed with the click of a button.

Open local files
JBrowse Web now has the ability for tracks to use files on your local hard drive. This is a great option if you want to visualize files you have locally without uploading them to a server. These files will need to be re-opened each time the app is opened or refreshed, but more robust handling of local files will be available when we release JBrowse Desktop.

Color by MM and MP/ML tags in BAM/CRAM
The MM and MP/ML tags can be used to color alignments tracks by either base modifications or by methylation. The modifications mode is exciting because it can show arbitrary DNA/RNA modifications, and the methylation mode uses specific CpG context to show both modified and unmodified CpGs.
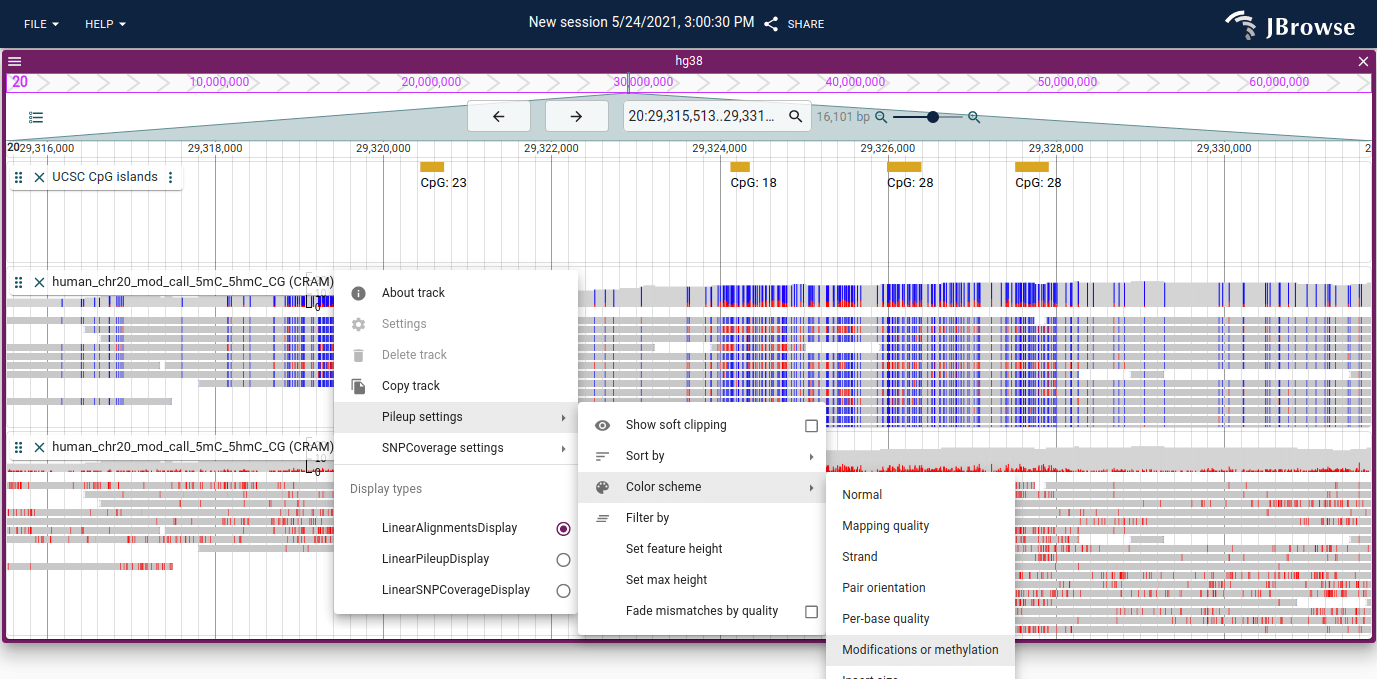
In this screenshot, the top alignments track is colored by methylation and the bottom alignments track is colored by base modification.
Downloads
To install JBrowse 2 for the web, you can download the link above, or you can use the JBrowse CLI to automatically download the latest version. See the JBrowse web quick start for more details.
1.3.0 (2021-05-24)
:rocket: Enhancement
- Other
core- #1982 Allow manually specifying adapter type if filename does not match expected pattern (@cmdcolin)
- #1975 Allow local files on the users computer to be opened as tracks in jbrowse-web (@cmdcolin)
- #1865 Show modified bases using MM and MP/ML tags in BAM/CRAM (@cmdcolin)
- #1984 Better feature details when there are short arrays of json supplied as feature data (@cmdcolin)
- #1931 Create in app graphical plugin store (@elliothershberg)
:bug: Bug Fix
core- #1985 Avoid error calculating UTR on features that have no exon subfeatures (@cmdcolin)
- #1954 Add more environments to configSchema create calls to fix ability to use custom jexl commands with main thread rendering (@cmdcolin)
- #1963 Fix ability to use DialogComponent (used for svg export, pileup sort, etc) on embedded components (@cmdcolin)
- #1945 Fix hic not being able to render due to incorrect lazy loading (@cmdcolin)
- Other
:memo: Documentation
- #2002 Add @jbrowse/img to homepage (@cmdcolin)
- #2007 Update docs for modifications/methylation coloring, plugin store, and the sequence panel in feature details (@cmdcolin)
- #1976 reorganize the demo page to emphasize the cancer sv demo more (@rbuels)
- #1952 Add demo for 1000 genomes extended trio dataset to website (@cmdcolin)
- #1862 Add example for using a build-time included plugin to storybook (@cmdcolin)
:house: Internal
- Other
core,development-tools
Committers: 4
- Colin Diesh (@cmdcolin)
- Elliot Hershberg (@elliothershberg)
- Garrett Stevens (@garrettjstevens)
- Robert Buels (@rbuels)